Wild Emmer – also called wild wheat – is an ancient tetraploid wheat (Triticum dicoccoides) from the Middle East, considered to be an ancestor of modern cultivated polyploid wheats, including Triticum aestivum (bread) and Triticum durum (pasta).
Wild Emmer itself is too low-yielding to be of use to farmers today, but it contains many of the characteristics that are being used by plant breeders to improve wheat.
More efficient than ever
A team of international scientists collaborated with Israeli genomic data company NRGene to develop a bioinformatics technology to fast-track the identification of the complex 14 chromosome sequence of Wild Emmer.
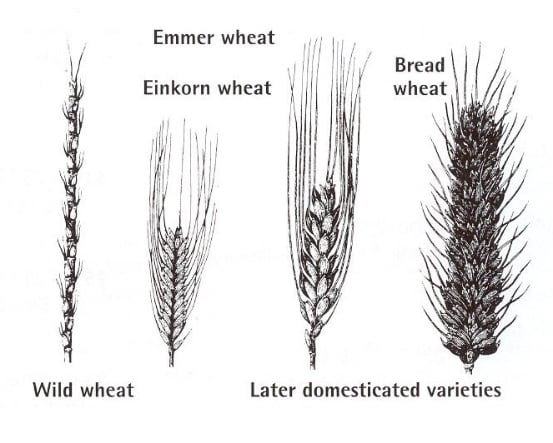
This ability now gives scientists the chance to examine wheat from before the origins of agriculture and compare it to modern wheat, thus cherry-picking the more advantageous genes and characteristics that were involved in domestication.
According to lead scientist Dr Assaf Distelfeld of Tel Aviv University’s School of Plant Sciences and Food Security and Institute for Cereal Crops Improvement, the new technology has provided scientists with tools to study crops directly and apply their discoveries more efficiently than ever before.
Food security revolution
Dr Curtis Pozniak from the University of Saskatchewan and chair of the Canadian Ministry of Agriculture Strategic Research Program, said:"Wheat accounts for almost 20% of the calories humans consume worldwide, so a strong focus on improving the yield and quality of wheat is essential for our future food supply,” said “This new resource allowed us to identify a number of other genes controlling main traits that were selected by early humans during wheat domestication and that served as foundation for developing modern wheat cultivars," said Dr Eduard Akhunov of Kansas State University.
“Wild Emmer is known as a source of novel variation that can help to improve the nutritional quality of grain as well as tolerance to diseases and water-limiting conditions."
Dr Zvi Peloeg of the Hebrew University of Jerusalem, Israel, added: "While many modern wheat cultivars are susceptible to water stress, Wild Emmer has undergone a long evolutionary history under the drought-prone Mediterranean climate. Thus, utilization of the wild genes in wheat breeding programs promotes producing more yield for less water."
Computing chromosomes
Dr Gil Ronen, NRGene’s CEO, said the wheat genome is four times the size of a human genome and is very large and complex.
Thanks to the company’s computational technology, though, the sequences of the 14 chromosomes of Wild Emmer wheat were easily collapsed into a refined order.
The technology has been successful in unveiling the much smaller human and barley genome sequences and has now paved the way to sequence large cereal genomes like durum wheat to understand how humanity transformed it into a high-yielding and high-quality crop, explained Dr Luigi Cattivelli, head of the CREA Research Centre for Genomics and Bioinformatics (Italy) and coordinator of the International Durum Wheat Genome Sequencing Consortium.
"This Wild Emmer wheat sequencing and approach is an invaluable contribution to the entire wheat community to improve and better understand nutritional mechanisms," said Dr Hikmet Budak, Montana Plant Science Endowed Chair at Montana State University.

